Addressing the Problem of Feature Selection Using Genetic Algorithms


The problem of feature selection
The explosion in the availability of big data has made complex prediction models a conspicuous reality of our times. Whether in banking, financial services and insurance, telecoms, manufacturing, or healthcare, predictive models are increasingly used to derive inferences from data.
Most of these models use a set of input variables, called features, to predict the output of a variable of interest. For example, the concentration of characteristic biomarkers in the blood can be used to predict the presence, absence or progress of certain diseases.
The available data can provide a large number of features, but generally, it’s preferable to use a small number of really relevant features in a model. This is because a model with more features has a greater complexity which leads to greater demand on computational resources and time to train the model. Therefore it is desirable to restrict the number of features in a predictive model. Choosing the subset of features that will result in a model with optimum performance is the problem of feature selection. This is essentially a problem of plenty.
To understand the magnitude of this problem, let us consider a typical scenario. If we have 100 features derived from a data set, (and this is fairly typical), and we wish to choose a subset of 10 features, then the number of possible combinations is more than a trillion. We are looking for a single combination of 10 features that gives us an optimally performing model. It is clearly not feasible to train the model on each of these trillion or more potential combinations to select the optimal one. That would be like looking for a needle in a haystack by examining every strand of hay!
Fortunately, there are intelligent methods apart from the one described above for searching through this massive feature space. One of these is the genetic algorithm (GA).
Genetic Algorithms
GA is a kind of evolutionary algorithm suited to solving problems with a large number of solutions where the best solution has to be found by searching the solution space. The algorithms draw ideas from evolutionary biology and genetics, in that they mimic processes of selection, cross-over, and mutation to get to optimal solutions. The processes we go on to describe below are analogs of similar processes in genetics. Before we continue to review an application of the genetic algorithm in another setting ( medical devices), let’s first walk through some of the terminology used.
Terminology
Each solution is likened to a “chromosome “
The set of possible solutions is regarded as a “population”.
The solutions are applied to a problem and for each solution a fitness value (a measure of how good the model is) is computed.
The solutions with the highest fitness values in the population are stochastically selected. These solutions are then used for “cross-over” to get the next generation of solutions. Crossover is the process of taking more than one parent solution and producing a child solution from them.
In order to maintain diversity in the population, “mutation“ of chromosomes is done with a low probability. In genetics “mutation” is randomly changing the state of the “gene” in the chromosome. In our application, genes in the chromosome are analogous to the components into which the solution is encoded, and “mutation” is randomly executing changes to the components to improve our chances of generating different solutions.
“Generation” refers to each iteration of the algorithm. The new generation of solutions are then applied to the problem and the process is repeated. The process is said to have converged when the highest fitness values among subsequent generations remain the same (within a certain tolerance).
The following figure is a schematic representing the essential elements of the algorithm.
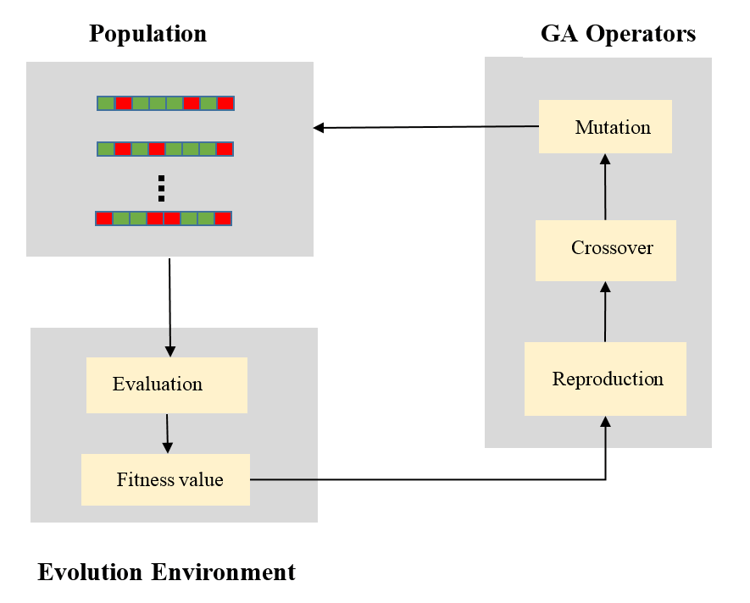
A schematic showing the components of a genetic algorithm.
[Ref: proteomicsnews.blogspot.in - 09-Nov-2016]
A Sample Use Case
Let us consider a sample problem in which we have data from a medical device that is used to monitor breathing. The data is measurement on several variables for subjects, and the classification problem is to determine whether the breathing action of a particular subject is normal or labored. The measurements are on data that are not easily mapped to visible attributes. We wish to select features that result in a model that performs optimally on this classification task.
A GA was run on this problem where we were looking for the best features for a classification model. The classification model was constructed on 10000 random training and test splits of data for each chromosome of a population in every generation. The fitness of the model was the median area under the ROC curve. The following graph illustrates the performance of the GA.
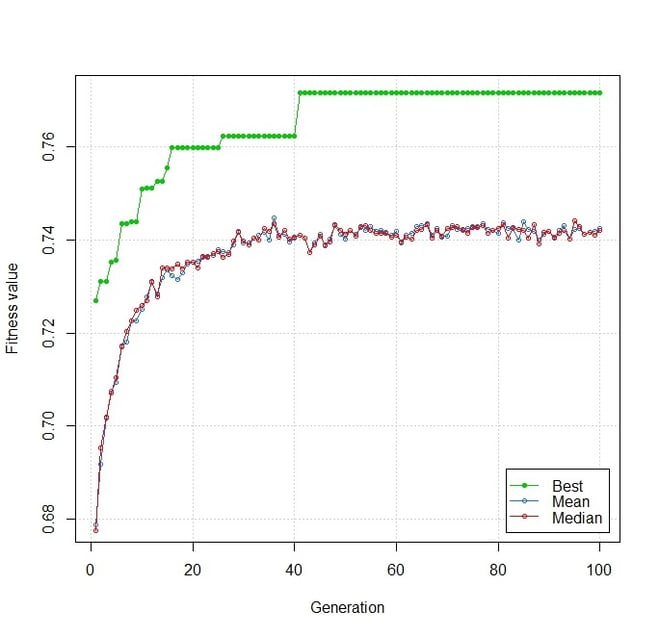
A plot of fitness values for chromosomes across generations.
Each generation had a population size of 100 chromosomes and the algorithm was run over 100 generations. As the number of generations increases, the fitness of the best chromosome in each generation, as indicated by the green line, increases. The mean fitness value of chromosomes in a generation, as shown by the blue line, also increases. This indicates that the candidates in subsequent generations have a better average fitness.
Beyond 50 generations, however, the mean fitness has saturated to a value of about 0.74. This means that the population of chromosomes seems to have reached a state where any further cross-over does not generate chromosomes with markedly better fitness. This could either mean that we have reached an optimal solution or that the population has lost genetic diversity. The fitness of the best solution obtained by the GA is about 0.77.
This was further tested by increasing the probability of mutation, to improve genetic diversity, as well as increasing the number of generations over which the algorithm was run. These changes, however, did not result in an improvement in fitness. This probably suggests that the optimal solution obtained is the best over most of the search space.
Conclusion
In the new era of big and complex data, conventional methods of solving search problems are sometimes ineffective. They may get trapped in local optima; the absence of a derivative may make them inapplicable, or they may be computationally too complex to solve in reasonable time and memory.
Genetic algorithms provide a highly effective solution to many such problems, and therefore their popularity and adoption are on the rise.
Cytel data scientists apply advanced statistical techniques including predictive modeling of biological processes and drug interactions to unlock the potential of big data. Our team supports biomarker discovery and diagnostic test development based on biomedical signals and images, and real-world evidence analysis.
To learn more about our data science services, click the button below to download the brochure.
About the Author
 Munshi Imran Hossain is a Software Affiliate at Cytel and currently works as a data scientist for strategic consulting assignments. He has 5 years of experience working in the areas of software development and data science. He holds an M. Tech in Biomedical Engineering, from IIT Bombay. His interests include processing and analysis of biomedical data. Outside of work, he enjoys reading.
Munshi Imran Hossain is a Software Affiliate at Cytel and currently works as a data scientist for strategic consulting assignments. He has 5 years of experience working in the areas of software development and data science. He holds an M. Tech in Biomedical Engineering, from IIT Bombay. His interests include processing and analysis of biomedical data. Outside of work, he enjoys reading.
Further reading
Blog: Removing Noise From Biomedical Signals
Blog: Signal Management Using R
Blog: Pattern Recognition and Big Data


