Harnessing the power of R API to extend software applications

 In the complex world of trial design and data analysis biostatisticians and data scientists need to ensure they are selecting and harnessing the best capabilities of the powerful software tools available to them. Particularly when non-standard approaches are required, this may mean using a combination of tools to come to the most appropriate solution for any task.
In the complex world of trial design and data analysis biostatisticians and data scientists need to ensure they are selecting and harnessing the best capabilities of the powerful software tools available to them. Particularly when non-standard approaches are required, this may mean using a combination of tools to come to the most appropriate solution for any task.
At the recent EARL conference, Cytel’s Aniruddha Deshmukh, Software Evangelist, discussed how R can be harnessed to extend and customize the powerful capabilities of East. Using R API , it’s possible to execute R code and manipulate R objects which are coded in other languages such as C/C++ and this approach has been used in East® to extend its features as well as customize simulations to meet any non-standard needs of a given clinical trial.
In this blog Aniruddha will take a techical look at R API and some of its key elements
What is R API?
R API is a collection of functions and data structures provided by R. It can be used in C/C++/Fortran code.
What does R API allow the user to do?
It allows the user to create and end an embedded R instance, as well as create and manipulate R variables from within your application. It’s possible, using R API to pass data back and forth between an application ( such as East) and R, as well as form and evaluate R expressions and commands and load and execute R scripts and functions.
A Technical Walkthrough
In order to use the R API in your code, you need to include the header file R.h and link your program with R.dll.
R.h is the main header file and it can be found in the folder ‘R_HOME\include’. It declares the API functions.
R.dll is the main DLL behind R that exports a large number of entry point functions. It can be found in the folder ‘R_HOME\bin\i386’ or ‘R_HOME\bin\x64’.
In addition to the functions, the R API also consists of an important C structure called SEXPREC. A pointer to SEXPREC is called SEXP. This is a complex structure that R uses to represent different object types including vectors, lists, data frames, function objects etc.
Several API functions take SEXP arguments and return SEXP.
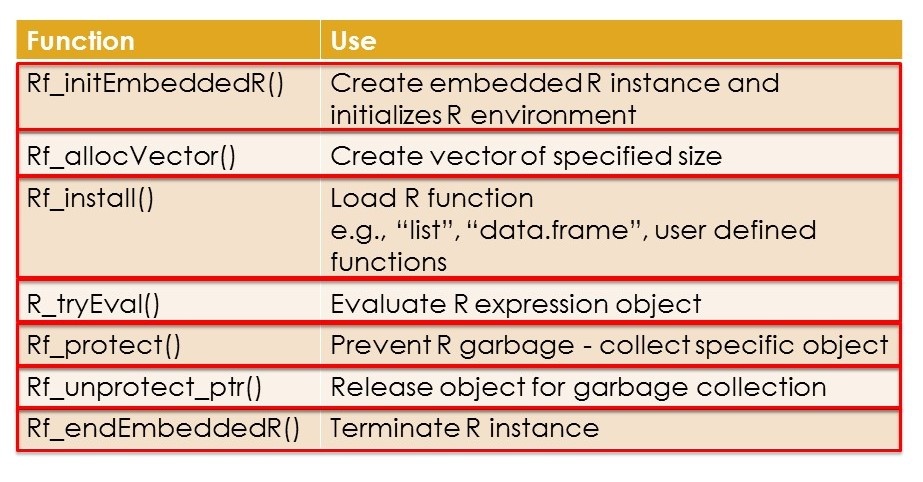
Below is a list of some useful API functions, with description of each function.
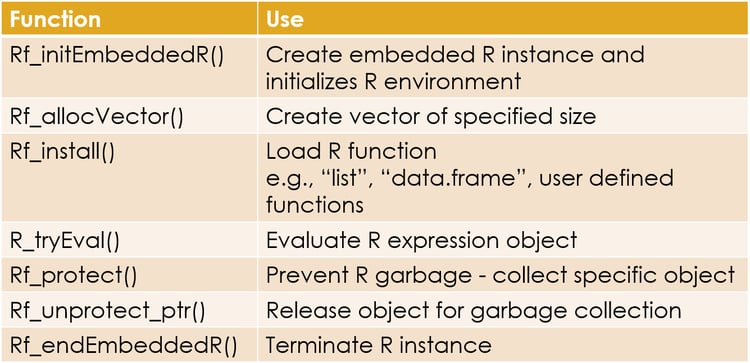
An important point to remember is that R uses garbage collection i.e. when R thinks that an object is no longer is use, it releases the memory allocated to this object and destroys it. This can be a problem for your program as R cannot know if you are using any R object inside your program. It may garbage collect this object and can cause your program to malfunction. To prevent this from happening, we use the API Rf_protect() and pass a SEXP pointer to the R object to it. This tells R to leave the object alone. When you are done using the object, you release the lock on it by calling the Rf_unprotect_ptr() function.
Now, let’s review some sample code. Consider the below R function. All it does is take an numeric argument, increment it and return it. Although, it’s very simple, it is sufficient to illustrate the key ideas.
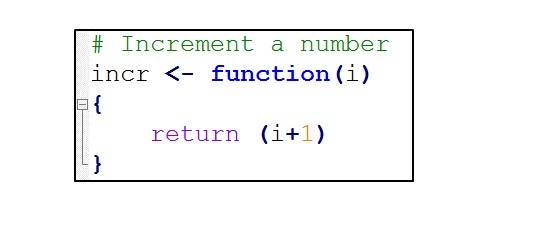

Now, suppose we want to execute this R function from our C code.
Step 1: We create an embedded instance of R using the API Rf_initEmbeddedR(). The argume “--silent” indicates we don’t want to see the R GUI, but execute R in the background.

Step 2: Next we create a vector of length 1 to hold the value of the numeric argument to be passed to the function. The type INTSXP indicates we want to create an integer vector. The API INTEGER() can be used to access the internal array storage of the vector.

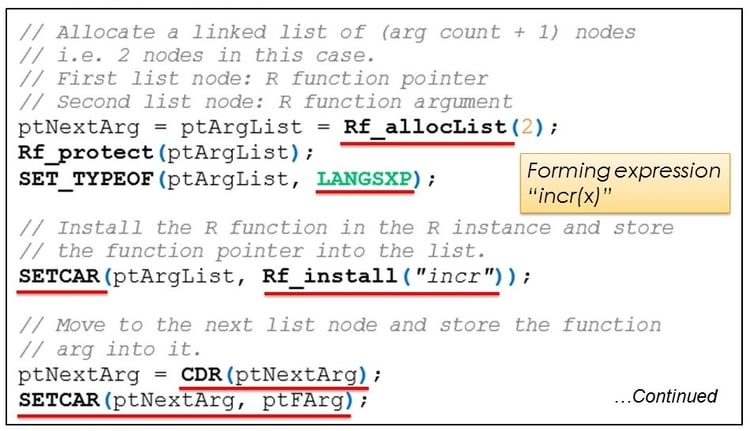
Step 3: We now want to create an R expression like “incr(x)” and evaluate it. To do this, we need to create a linked list of size 2. This is done using the API Rf_allocList().
The first node in this list holds a pointer to the R function we want to execute. We get a function pointer by executing the function Rf_install() and passing the R function name to it. This loads the R function in the global environment and returns a pointer to it.
The function SETCAR() is used to store the function pointer into the first node of the linked list.
We move to the second node by calling the function CDR() and then call SETCAR() again to store the function argument into the second node of the linked list.
The fully created linked list is now equivalent to the R expression “incr(x)”. It is ready to be evaluated.
Final step
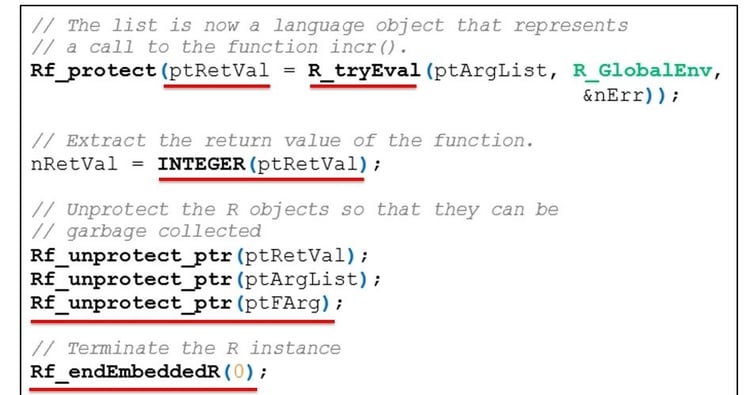
To evaluate the R expression, we call the function R_tryEval() and pass the linked list pointer to it. This function evaluates the expression and returns its value. If the evaluation fails, the argument nErr holds the error code.
We use the INTEGER() function to extract the value returned by the R function. We now have the function’s return value available in a format that our C code can understand.
Finally we release all R objects for garbage collection by unprotecting them and then terminate the embedded R instance.
Summary
The same approach can work while creating and manipulating lists and data frames or executing complex R functions and scripts. It also enables the loading of R packages and executing the functions within them. If used appropriately it can help customize a given application. However, caution should always be taken as always with using externally generated R user code.
Want to review this article offline? Click the link below to download a pdf copy.
Related reading:
The Rise of R- should SAS Programmers get up to speed?


